2009-2013
My self-initiated research began in the fall of 2009, and over the next 4 years I developed several devices, software, and an iPhone app.
I started meditating and immediately began looking for an objective way to track my progress. In December, I built my first device for amplifying the heartbeat intervals using an off-the-shelf photoplethysmography finger-tip sensor (PPG) that was then connected to a laptop and analyzed with code I wrote in LabVIEW.
Not content to be tethered to my laptop, and annoyed by how noisy PPG is, I soon built a biofeedback device using a microcontroller (Logomatic v2), screen (ST7565), and a Polar receiver (RMCM01) to wirelessly receive heartbeats from a Polar HR T31 strap (which sends a 5 kHz radio pulse each beat).
2010 sleep interest
That summer, I discovered that if I wore the strap while sleeping, I observed clear transitions and cycles. At this point I knew nothing about sleep stages or the research on them (and sleep wasn’t in the zeitgeist yet). However, this marked the beginning of my interest in sleep stages, and in the idea of measuring them with the heart.
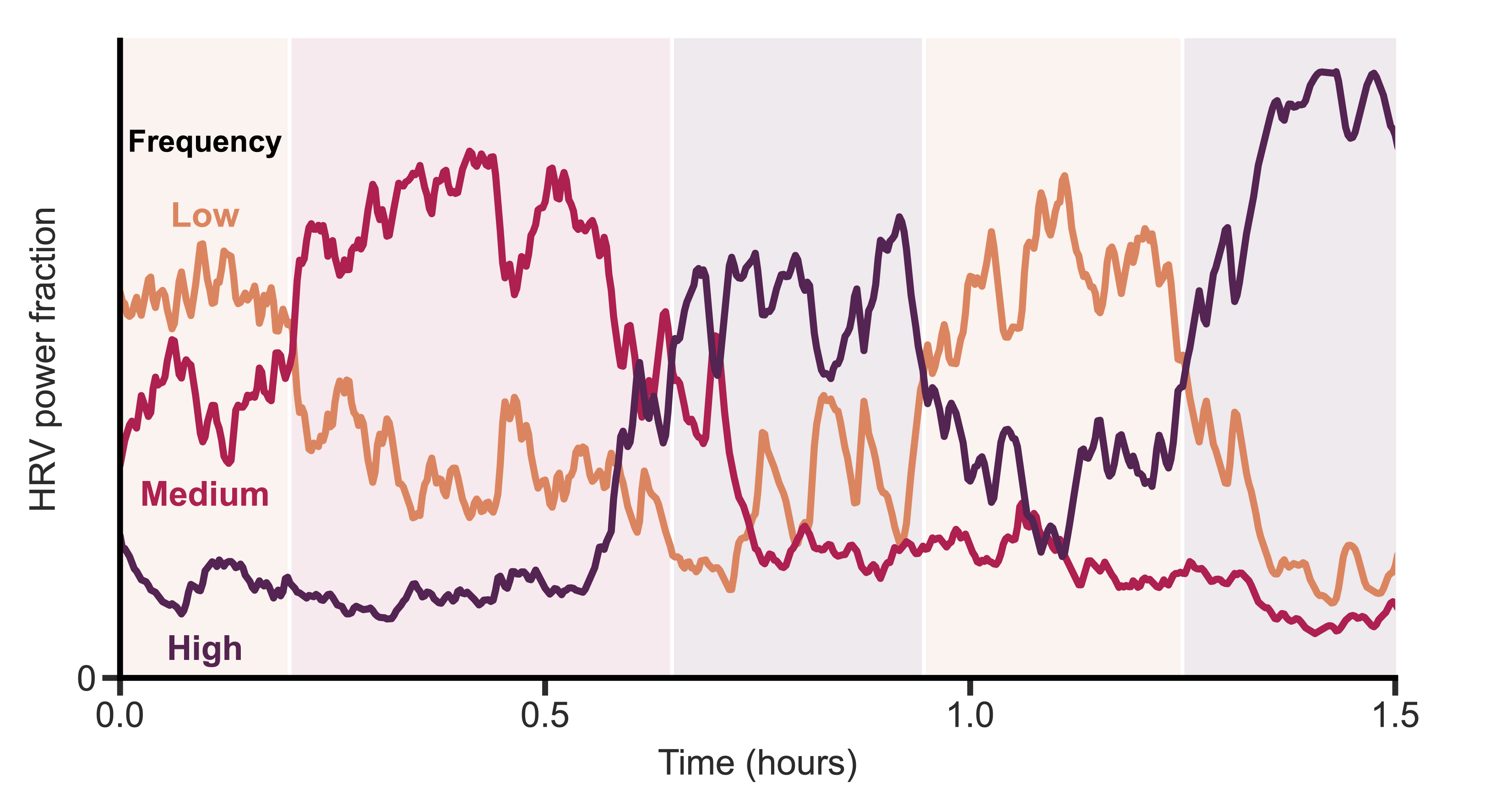
I also began recruiting beta testers among my friends and coworkers. I wanted to see what the variations in data looked like, and how robust the device and algorithms were. At the time, I only had a single device, so this meant loaning it out each time.
Not content with the low-resolution monochromatic screen, and the unnecessary bulk of my pocketable (but still too-large) device, I decided to build an iPhone app.
-
Plan A: To keep things simple, I stuck with the 5 kHz strap+receiver. I designed my first manufactured PCB to attach to the iPhone’s 30-pin dock connector.
I had thought about using the connector’s audio input to receive the voltage blips from the receiver IC directly. However, I wasn’t yet confident that I wouldn’t miss any of the blips. So, I had a real-time microcontroller act as the middleman.
iPhone heartbeat receiver dongle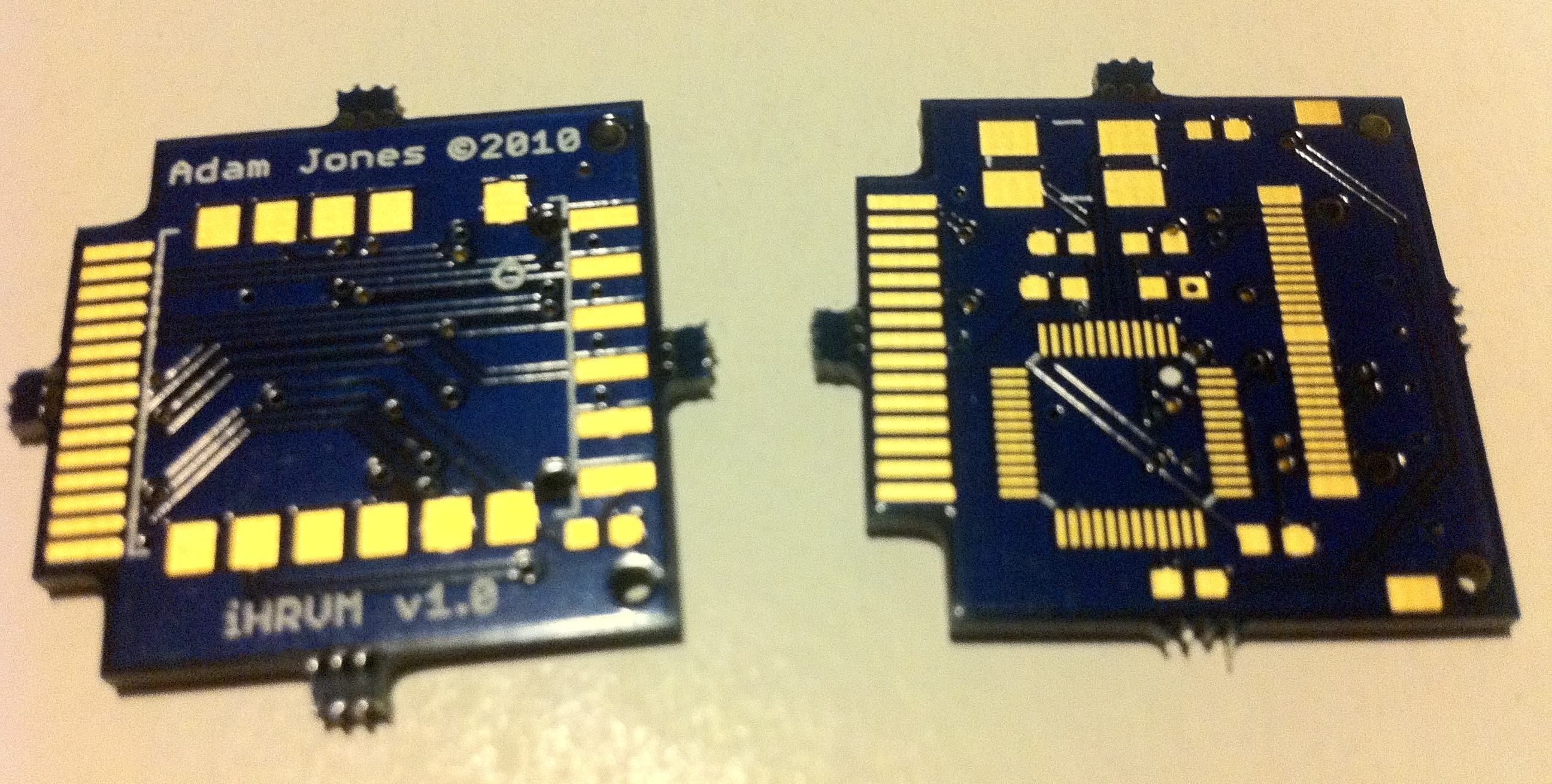
Unfortunately, when I received the PCBs, I realized they were too thick for the connector’s legs 😭 (which were meant to straddle the top and bottom). However, in the weeks it took between uploading the design files and the finished board arriving, I had set my sights on an even better idea…
-
Plan B: The iPhone 4 (“you’re holding it wrong”) was released that year and had Bluetooth 2.1. I also stumbled upon one of the first Bluetooth HR straps, the Zephyr HxM Bluetooth. The final missing piece was a developer that created a BT API for jailbroken phones (I can’t seem to recall the name of the library right now), since Apple didn’t yet give developers an API to access BT.
A year or so later I found two companies selling headphone dongles that received the 5 kHz pulse directly. So, even though the BLE strap was my preferred input, I designed my app to allow the user to select either a BLE strap or a “traditional” 5 kHz strap with a separately purchased headphone dongle.
Over the next several years, I quickly learned the ropes of app development and programming best practices that I hadn’t yet learned (having taught myself GW-BASIC around 1989 and learning a dozen more languages along the way).
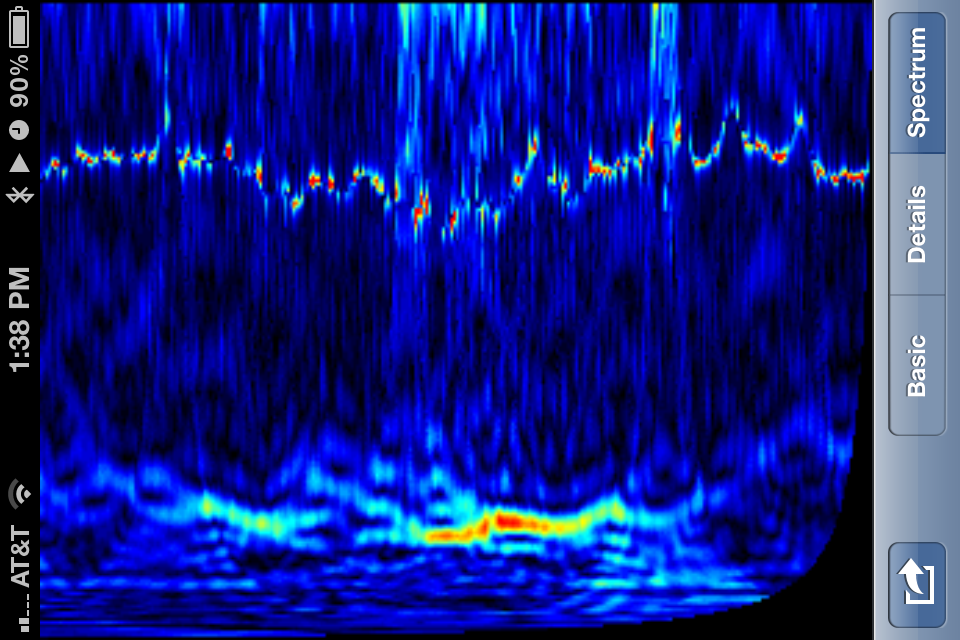
I began attending local iPhone developer meetups and recruiting more beta testers. Now the testers just needed an iPhone (or for the one Android user, I mailed one), and I could loan them one of several straps.
2014-2018
When I began working on a BS in psychology at the University of Houston, this research then expanded into a collaboration with Dr. Bhavin R. Sheth. We decided to tackle trying to score sleep using ECG from a small dataset (n=63) from the Veterans’ Affairs (VA).
While I certainly learned a lot of analysis techniques during my BS and MS in mechanical engineering, I now had to teach myself machine learning and robust statistics.
At the time, I was converting the ECG recordings into RR intervals. This is when I found out that algorithms for doing this are… to be kind, not very robust. Every single algorithm and tool I could get my hands on required manual intervention and annotation on all but the most clean and pristine data; none of them handled noise gracefully. Thus began a side project of building a much more robust heartbeat (R wave) detector.
Developing my new robust heartbeat algorithm took on even more importance when I stumbled upon the National Sleep Research Resource (NSSR) and the Sleep Heart Health Study (SHHS) dataset, which took us from 63 recordings to over 8,000. There was no way on earth I could babysit even the best of the existing algorithms on every single recording.
2015 SfN (k=0.280)
I gave my first talk at the 2015 Society for Neuroscience’s (SfN) annual meeting in Chicago. At the time, I was using combinations of “traditional” machine learning techniques on tons of hand-crafted features (from the RR itself, the spectrum of the RR, etc.), but the results were not great. The Cohen’s kappa was 0.28—pretty abysmal. I also presented a new parametric non-sinusoidal function that I found better matched the RR intervals during respiratory sinus arrhythmia (RSA) than a simple sine wave.
However, a few months after my November 2015 talk, I gave up on RR intervals. And, much to the chagrin of my collaborator, I began throwing neural networks (NNs) at the problem (In 2007, I had explored using NNs during my MS as a way to find nonlinear functions to fit the very nonlinear data from my experiments, long before the deep learning breakthrough in 2012). Given that this was temporal data, I originally started with LSTM- and GRU-based architectures.
2017 SfN (k=0.530)
I presented these findings at the 2017 SfN meeting in D.C. on a “dynamic” poster (i.e., a large TV). The results were now a quite respectable Cohen’s kappa of 0.530—better than state-of-the-art on 5-stage scoring for “EEG-less” methods (any method for sleep staging that makes no use of brain, i.e., EEG data). For reference, the current state-of-the-art on 5-stage scoring was k=0.510 from Sady et al, 2013 [1].
However, I wasn’t content with just state-of-the-art for EEG-less methods. My aim was now firmly fixed on matching the performance of human-scored polysomnography (PSG, or a traditional “sleep study”).
I needed a bigger (and better) model, and to train that, I also needed more data. The first thing to go was the RNN architecture. I began experimenting with a new “backbone”, the Temporal Convolution Network (TCN), which made the network completely feed-forward and faster to train.
That summer I began lab rotations for my neuroscience PhD at the University of Southern California.
2018 SfN (k=0.710)
I gave my second talk at the 2018 SfN meeting in San Diego, where I demonstrated that we had reached a Cohen’s kappa of 0.710 on 5-stage scoring. We were now significantly better than state-of-the-art, and finally within the range of expert human-scored PSG.
It was then that I remembered my original HRV device, and felt I needed to begin experimenting on myself. Finding, once again, that there was no reasonably-priced device (narrator: there was, but it was only pointed out to him years later), I started designing one. I was going to make it wireless, for comfort, and attach to the “bare” Polar strap (they have snaps for attaching the wireless modules, so that the strap can be washed).
I ordered the parts, and started experimenting with various techniques of recording a clean signal without a separate ground electrode. However, my required coursework and lab rotations just took up most of my time, and I was feeling impatient. So, I did what I thought was the next-best thing: I ordered the recently-released 2nd generation Oura Ring, in the hopes that it might be good enough.
2019-2021
So, the projects (both the model and the hardware) were on hold while I worked on research for my PhD with Dr. Laurent Itti. I think there was a pandemic somewhere in here, and time lost all meaning. However, every few months I would occasionally experiment with different aspects of the model and training.
In 2020 a new, published, state-of-the-art threshold was reached for EEG-less methods, k=0.585 from Sun et al. [2]. However, unbeknownst to anyone that didn’t attend my 2018 SfN talk, this was significantly below the k=0.710 I had already presented.
2022-2024
After a 3-year hiatus, I began working on the research again. My model had been collecting dust—the world none the wiser and unable to use it. So, this time, I had set my sights on publishing the model and releasing the code to the world.
For my PhD qualifying exam in January, I switched my final PhD project to “finishing up” the not-yet-published sleep staging model as well as the extensions I had long known were in the pipeline (see Future Plans below).
In the midst of greatly expanding the training data (making sure to never test on any subject I had ever trained any previous model on), I found a glaring issue with one of the NSRR datasets. So, since I had to replace thousands of recordings, I decided to also target a smoother age distribution.
While putting the finishing touches on the first paper, I realized there was a bigger story that could be told. The additional analyses, including those suggested from an invited talk I gave, delayed the paper’s submission by several months.
However, by this time, I had already started to work on the next phase of the research in parallel. Unfortunately, the details of this phase will have to remain under wraps until the next paper is published.
In November, we submitted the paper to Computers in Biology and Medicine (CIBM).
2023 PhD defense
For those in attendance, they got to see the main findings in the first paper, as well as the aforementioned second phase that will make up the next paper or two.
2024 CIBM (k=0.725)
To address the biggest concern from the initial round of reviews, I taught myself about meta-analyses and non-inferiority testing. And, after another two rounds of reviews, the paper was accepted by CIBM on April 28th.
Future Plans
Models and code
The most immediate plans I have for the research are the following:
- To finish converting the data preprocessing pipeline from MATLAB to Python, so that others can more easily (and freely) use it.
- To test two of the commercial sensors (Blog update: I’ve begun testing them).
- To write and release a simple 100% free iPhone app for recording and saving the data from those sensors.
Research
During my PhD defense I presented additional findings that aren’t in the current paper. However, I put my dissertation under embargo to give me some time to get the next paper or two out. These additional findings are also likely to be surprising.
Call for Input
Since the first paper was published, I’ve been privately contacted by numerous researchers and clinicians around the world on using and extending the model to make it even more useful to them and sleep medicine in general. If you’re involved in sleep medicine, please reach out to me, as I’m trying to get as many ideas and perspectives as possible on where to extend this research to have the greatest benefit for human wellbeing.
References:
Originally posted on cardiosomnography.com.
]]>